For the full list of publications, scroll down.
Featured work
Statistical finite elements for misspecified models
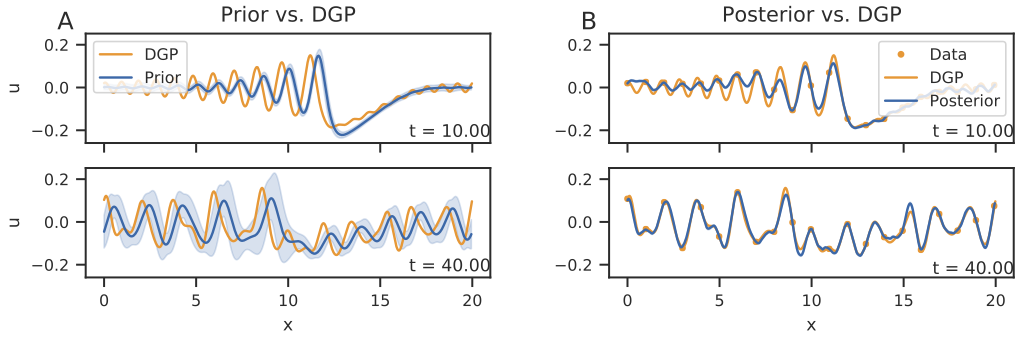
We develop the statistical FEM methodology in the context of nonlinear internal waves (solitons), using the Korteweg-de Vries equation.
Connor Duffin, Edward Cripps, Thomas Stemler, and Mark Girolami
Proceedings of the National Academy of Sciences
University of Cambridge coverage
See the research section for more details.
Riemann manifold Langevin and Hamiltonian Monte Carlo methods
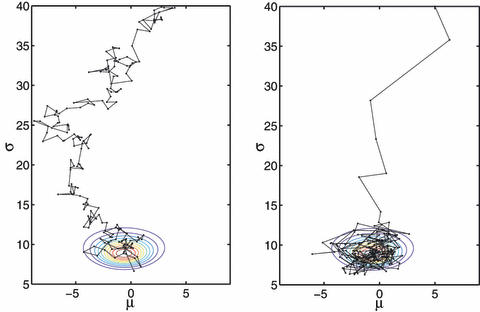
This paper proposes Metropolis adjusted Langevin and Hamiltonian Monte Carlo sampling methods defined on the Riemann manifold to resolve the shortcomings of existing Monte Carlo algorithms when sampling from target densities that may be high dimensional and exhibit strong correlations.
Girolami, M. and Calderhead, B
Journal of the Royal Statistical Society: Series B (Statistical Methodology)
See the research section for more details.
All publications
Zachos, Ioannis, Damoulas, Theodoros, & Girolami, Mark. Table inference for combinatorial origin-destination choices in agent-based population synthesis. Stat, 13(1), e656, mar 2024. URL: https://doi.org/10.1002/sta4.656, doi:10.1002/sta4.656.
Connor Duffin, Edward Cripps, Thomas Stemler, and Mark Girolami. Low-rank statistical finite elements for scalable model-data synthesis. Journal of Computational Physics, 463:111261, aug 2022. URL: https://doi.org/10.1016%2Fj.jcp.2022.111261, doi:10.1016/j.jcp.2022.111261.
Jan Povala, Ieva Kazlauskaite, Eky Febrianto, Fehmi Cirak, and Mark Girolami. Variational bayesian approximation of inverse problems using sparse precision matrices. Computer Methods in Applied Mechanics and Engineering, 393:114712, apr 2022. URL: https://doi.org/10.1016%2Fj.cma.2022.114712, doi:10.1016/j.cma.2022.114712.
P. Gardner, L.A. Bull, J. Gosliga, J. Poole, N. Dervilis, and K. Worden. A population-based SHM methodology for heterogeneous structures: transferring damage localisation knowledge between different aircraft wings. Mechanical Systems and Signal Processing, 172:108918, jun 2022. URL: https://doi.org/10.1016%2Fj.ymssp.2022.108918, doi:10.1016/j.ymssp.2022.108918.
Lawrence A. Bull, Nikolaos Dervilis, Keith Worden, Elizabeth J. Cross, and Timothy J. Rogers. A sampling-based approach for information-theoretic inspection management. Proceedings of the Royal Society A: Mathematical, Physical and Engineering Sciences, jun 2022. URL: https://doi.org/10.1098%2Frspa.2021.0790, doi:10.1098/rspa.2021.0790.
L.A. Bull, P.A. Gardner, T.J. Rogers, N. Dervilis, E.J. Cross, E. Papatheou, A.E. Maguire, C. Campos, and K. Worden. Bayesian modelling of multivalued power curves from an operational wind farm. Mechanical Systems and Signal Processing, 169:108530, apr 2022. URL: https://doi.org/10.1016%2Fj.ymssp.2021.108530, doi:10.1016/j.ymssp.2021.108530.
A.J. Hughes, L.A. Bull, P. Gardner, R.J. Barthorpe, N. Dervilis, and K. Worden. On risk-based active learning for structural health monitoring. Mechanical Systems and Signal Processing, 167:108569, mar
Domenic Di Francesco, Mark Girolami, Andrew B. Duncan, and Marios Chryssanthopoulos. A probabilistic model for quantifying uncertainty in the failure assessment diagram. Structural Safety, 99:102262, nov
H. Rappel, M. Girolami, and L. A. A. Beex. Intercorrelated random fields with bounds and the bayesian identification of their parameters: application to linear elastic struts and fibers. International Journal for Numerical Methods in Engineering, mar 2022. URL: https://doi.org/10.1002%2Fnme.6974, doi:10.1002/nme.6974.
Patricio Peralta, Rafael O. Ruiz, Hussein Rappel, and Stéphane P.A. Bordas. Electromechanical properties identification for groups of piezoelectric energy harvester based on bayesian inference. Mechanical Systems and Signal Processing, 162:108034, jan 2022. URL: https://doi.org/10.1016%2Fj.ymssp.2021.108034, doi:10.1016/j.ymssp.2021.108034.
B. Boys, T.J. Dodwell, M. Hobbs, and M. Girolami. - a high performance OpenCL peridynamics package. Computer Methods in Applied Mechanics and Engineering, 386:114085, dec 2021. URL: https://doi.org/10.1016%2Fj.cma.2021.114085, doi:10.1016/j.cma.2021.114085.
Toni Karvonen, Chris J. Oates, and Mark Girolami. Integration in reproducing kernel hilbert spaces of gaussian kernels. Mathematics of Computation, pages 1, jun 2021. URL: https://doi.org/10.1090%2Fmcom%2F3659, doi:10.1090/mcom/3659.
Steven A. Niederer, Michael S. Sacks, Mark Girolami, and Karen Willcox. Scaling digital twins from the artisanal to the industrial. Nature Computational Science, 1(5):313–320, may 2021. URL: https://doi.org/10.1038%2Fs43588-021-00072-5, doi:10.1038/s43588-021-00072-5.
Connor Duffin, Edward Cripps, Thomas Stemler, and Mark Girolami. Statistical finite elements for misspecified models. Proceedings of the National Academy of Sciences, dec 2020. URL: https://doi.org/10.1073%2Fpnas.2015006118, doi:10.1073/pnas.2015006118.
Rebecca Ward, Ruchi Choudhary, Alastair Gregory, Melanie Jans-Singh, and Mark Girolami. Continuous calibration of a digital twin: comparison of particle filter and bayesian calibration approaches. Data-Centric Engineering, 2021. URL: https://doi.org/10.1017%2Fdce.2021.12, doi:10.1017/dce.2021.12.
Ömer Deniz Akyildiz and Joaquı́n Mı́guez. Convergence rates for optimised adaptive importance samplers. Statistics and Computing, jan 2021. URL: https://doi.org/10.1007%2Fs11222-020-09983-1, doi:10.1007/s11222-020-09983-1.
P Gardner, LA Bull, N Dervilis, and K Worden. Overcoming the problem of repair in structural health monitoring: metric-informed transfer learning. Journal of Sound and Vibration, pages 116245–116245, Jun 2021. doi:10.1016/j.jsv.2021.116245.
Lawrence A. Bull, Paul Gardner, Timothy J. Rogers, Elizabeth J. Cross, Nikolaos Dervilis, and Keith Worden. Probabilistic inference for structural health monitoring: new modes of learning from data. ASCE-ASME Journal of Risk and Uncertainty in Engineering Systems, Part A: Civil Engineering, 7(1):03120003, mar 2021. URL: https://doi.org/10.1061%2Fajrua6.0001106, doi:10.1061/ajrua6.0001106.
LA Bull, P Gardner, TJ Rogers, EJ Cross, N Dervilis, and K Worden. New modes of inference for probabilistic shm. In Lecture Notes in Civil Engineering, pages 415–426. Springer International Publishing, 2021. doi:10.1007/978-3-030-64908-1_39.
J Gosliga, P Gardner, LA Bull, N Dervilis, and K Worden. Towards population-based structural health monitoring, part ii: heterogeneous populations and structures as graphs. In Topics in Modal Analysis & Testing, Volume 8, pages 177–187. Springer International Publishing, 2021. doi:10.1007/978-3-030-47717-2_17.
Zheng Zhao, Toni Karvonen, Roland Hostettler, and Simo Sarkka. Taylor moment expansion for continuous-discrete gaussian filtering. IEEE Transactions on Automatic Control, 66(9):4460–4467, sep 2021. URL: https://doi.org/10.1109%2Ftac.2020.3047367, doi:10.1109/tac.2020.3047367.
Toni Karvonen, Simo Särkkä, and Ken’ichiro Tanaka. Kernel-based interpolation at approximate fekete points. Numerical Algorithms, 87(1):445–468, jul 2020. URL: https://doi.org/10.1007%2Fs11075-020-00973-y, doi:10.1007/s11075-020-00973-y.
Toni Karvonen, Simo Särkkä, and Ken’ichiro Tanaka. Correction to: kernel-based interpolation at approximate fekete points. Numerical Algorithms, dec 2020. URL: https://doi.org/10.1007%2Fs11075-020-01034-0, doi:10.1007/s11075-020-01034-0.
Jakub Pruher, Toni Karvonen, Chris J. Oates, Ondrej Straka, and Simo Sarkka. Improved calibration of numerical integration error in sigma-point filters. IEEE Transactions on Automatic Control, 66(3):1286–1292, mar 2021. URL: https://doi.org/10.1109%2Ftac.2020.2991698, doi:10.1109/tac.2020.2991698.
Onur Teymur, Christopher N. Foley, Philip G. Breen, Toni Karvonen, and Chris J. Oates. Black box probabilistic numerics. In M. Ranzato, A. Beygelzimer, Y. Dauphin, P.S. Liang, and J. Wortman Vaughan, editors, Advances in Neural Information Processing Systems 34 pre-proceedings (NeurIPS 2021), volume 34, 23452—–23464. 2021. Conference on Neural Information Processing System, NeurIPS 2021 ; Conference date: 06-12-2021 Through 14-12-2021.
Toni Karvonen, Simo Sarkka, and Christos Merkatas. Gaussian approximations of sdes in metropolis-adjusted langevin algorithms. In 31st IEEE International Workshop on Machine Learning for Signal Processing. 2021. doi:10.1109/MLSP52302.2021.9596301.
Toni Karvonen, Jakub Prüher, Ondřej Straka, Chris Oates, and Simo Sarkka. Improved calibration of numerical integration error in sigma-point filters. IEEE Transactions on Automatic Control, 66(3):1286–1292, 2021.
Toni Karvonen, Chris Oates, and Mark A. Girolami. Integration in reproducing kernel hilbert spaces of gaussian kernels. Mathematics of Computation, 90(331):2209–2233, 2021.
Toni Karvonen, Simo Sarkka, and Ken’ichiro Tanaka. Kernel-based interpolation at approximate fekete points. Numerical Algorithms, 87(1):445–468, 2021.
Toni Karvonen, Gabriele Santin, and Bernard Haasdonk. Sampling based approximation of linear functionals in reproducing kernel hilbert spaces. BIT Numerical Mathematics, 2021.
Toni Karvonen, Leah South, Chris Nemeth, Mark A. Girolami, and Chris Oates. Semi-exact control functionals from sard’s method. Biometrika, 2021.
Toni Karvonen, Zheng Zhao, Roland Hostettler, and Simo Sarkka. Taylor moment expansion for continuous-discrete gaussian filtering and smoothing. IEEE Transactions on Automatic Control, 66(9):4460–4467, 2021.
Domenic Di Francesco, Marios Chryssanthopoulos, Michael Havbro Faber, and Ujjwal Bharadwaj. Decision-theoretic inspection planning using imperfect and incomplete data. Data-Centric Engineering, 2021. URL: https://doi.org/10.1017%2Fdce.2021.18, doi:10.1017/dce.2021.18.
Maharshi Dhada, Mark Girolami, and Ajith Kumar Parlikad. Anomaly detection in a fleet of industrial assets with hierarchical statistical modeling. Data-Centric Engineering, 2020. URL: https://doi.org/10.1017%2Fdce.2020.19, doi:10.1017/dce.2020.19.
J. Povala, S. Virtanen, and M. Girolami. Burglary in london: insights from statistical heterogeneous spatial point processes. Journal of the Royal Statistical Society. Series C: Applied Statistics, 69(5):1067–1090, 2020.
Rafael Sacks, Ioannis Brilakis, Ergo Pikas, Haiyan Sally Xie, and Mark Girolami. Construction with digital twin information systems. Data-Centric Engineering, 2020. URL: https://doi.org/10.1017%2Fdce.2020.16, doi:10.1017/dce.2020.16.
Melanie Jans-Singh, Kathryn Leeming, Ruchi Choudhary, and Mark Girolami. Digital twin of an urban-integrated hydroponic farm. Data-Centric Engineering, 2020. URL: https://doi.org/10.1017%2Fdce.2020.21, doi:10.1017/dce.2020.21.
C.Y. Wong, P. Seshadri, G.T. Parks, and M. Girolami. Embedded ridge approximations. Computer Methods in Applied Mechanics and Engineering, 2020.
K. Menberg, A. Bidarmaghz, A. Gregory, R. Choudhary, and M. Girolami. Multi-fidelity approach to bayesian parameter estimation in subsurface heat and fluid transport models. Science of the Total Environment, 2020.
K. Monterrubio-Gómez, L. Roininen, S. Wade, T. Damoulas, and M. Girolami. Posterior inference for sparse hierarchical non-stationary models. Computational Statistics and Data Analysis, 2020.
S.-C. Kuok, K.-V. Yuen, S. Roberts, and M.A. Girolami. Propagative broad learning for nonparametric modeling of ambient effects on structural health indicators. Structural Health Monitoring, 2020.
Ömer Deniz Akyildiz, Dan Crisan, and Joaquı́n Mı́guez. Parallel sequential monte carlo for stochastic gradient-free nonconvex optimization. Statistics and Computing, 30(6):1645–1663, jul 2020. URL: https://doi.org/10.1007%2Fs11222-020-09964-4, doi:10.1007/s11222-020-09964-4.
Ömer Deniz Akyildiz and Joaquı́n Mı́guez. Nudging the particle filter. Statistics and Computing, 30(2):305–330, jul 2019. URL: https://doi.org/10.1007%2Fs11222-019-09884-y, doi:10.1007/s11222-019-09884-y.
Ayman Boustati, Omer Deniz Akyildiz, Theodoros Damoulas, and Adam Johansen. Generalised bayesian filtering via sequential monte carlo. Advances in Neural Information Processing Systems, 2020.
Lorenz Richter, Ayman Boustati, Nikolas Nüsken, Francisco Ruiz, and Omer Deniz Akyildiz. Vargrad: a low-variance gradient estimator for variational inference. Advances in Neural Information Processing Systems, 2020.
J Gosliga, PA Gardner, LA Bull, N Dervilis, and K Worden. Towards population-based structural health monitoring, part iii: graphs, networks and communities. In Topics in Modal Analysis & Testing. Proceedings of the 38th IMAC, A Conference and Exposition on Structural Dynamics 2020, volume 8, 255–267. Houston, TX, USA, Springer, Oct 2020. doi:10.1007/978-3-030-47717-2_26.
L.A. Bull, K. Worden, and N. Dervilis. Towards semi-supervised and probabilistic classification in structural health monitoring. Mechanical Systems and Signal Processing, 140:106653, jun 2020. URL: https://doi.org/10.1016%2Fj.ymssp.2020.106653, doi:10.1016/j.ymssp.2020.106653.
R Fuentes, EJ Cross, PA Gardner, LA Bull, TJ Rogers, RJ Barthorpe, H Shi, N Dervilis, CR Farrar, and K Worden. Structural health monitoring and damage identification. In Handbook of Experimental Structural Dynamics, pages 1–72. Jan 2020. doi:10.1007/978-1-4939-6503-8_23-1.
LA Bull, K Worden, TJ Rogers, EJ Cross, and N Dervilis. Investigating engineering data by probabilistic measures. In Special Topics in Structural Dynamics & Experimental Techniques, Volume 5, pages 77–81. Springer International Publishing, 2020. doi:10.1007/978-3-030-12243-0_12.
P Gardner, LA Bull, N Dervilis, and K Worden. Kernelised bayesian transfer learning for population-based structural health monitoring. In Model Validation and Uncertainty Quantification, Volume 3, pages 209–215. Springer International Publishing, 2020. doi:10.1007/978-3-030-47638-0_22.
LA Bull, PA Gardner, J Gosliga, N Dervilis, E Papatheou, AE Maguire, C Campos, TJ Rogers, EJ Cross, and K Worden. Towards population-based structural health monitoring, part i: homogeneous populations and forms. In Model Validation and Uncertainty Quantification, Volume 3, pages 287–302. Springer International Publishing, 2020. doi:10.1007/978-3-030-47638-0_32.
P Gardner, LA Bull, J Gosliga, N Dervilis, and K Worden. Towards population-based structural health monitoring, part iv: heterogeneous populations, transfer and mapping. In Model Validation and Uncertainty Quantification, Volume 3, pages 187–199. Springer International Publishing, 2020. doi:10.1007/978-3-030-47638-0_20.
Toni Karvonen and Simo Särkkä. Worst-case optimal approximation with increasingly flat gaussian kernels. Advances in Computational Mathematics, mar 2020. URL: https://doi.org/10.1007%2Fs10444-020-09767-1, doi:10.1007/s10444-020-09767-1.
Toni Karvonen, Simo Sarkka, George Wynne, Filip Tronarp, and Chris Oates. Maximum likelihood estimation and uncertainty quantification for gaussian process approximation of deterministic functions. SIAM/ASA journal on uncertainty quantification, 8(3):926–958, 2020.
Toni Karvonen, Silvère Bonnabel, Eric Moulines, and Simo Sarkka. On stability of a class of filters for nonlinear stochastic systems read more: https://epubs.siam.org/doi/10.1137/19m1285974. SIAM Journal on Control and Optimization, 58(4):2023–2049, 2020.
Toni Karvonen and Simo Sarkka. Worst-case optimal approximation with increasingly flat gaussian kernels. Advances in Computational Mathematics, 2020.
Domenic Di Francesco, Marios Chryssanthopoulos, Michael Havbro Faber, and Ujjwal Bharadwaj. Consistent and coherent treatment of uncertainties and dependencies in fatigue crack growth calculations using multi-level bayesian models. Reliability Engineering &$\mathsemicolon$ System Safety, 204:107117, dec 2020. URL: https://doi.org/10.1016%2Fj.ress.2020.107117, doi:10.1016/j.ress.2020.107117.
Ivan Ustyuzhaninov*, Ieva Kazlauskaite*, Markus Kaiser, Erik Bodin, Neill Campbell, and Carl Henrik Ek. Compositional uncertainty in deep gaussian processes. In Conference on Uncertainty in Artificial Intelligence, 480–489. PMLR, 2020.
Ivan Ustyuzhaninov*, Ieva Kazlauskaite*, Carl Henrik Ek, and Neill DF Campbell. Monotonic gaussian process flow. In International Conference on Artificial Intelligence and Statistics (AISTATS 2020). arXiv preprint arXiv:1905.12930, 2020.
H. Rappel, L.A.A. Beex, J.S. Hale, L. Noels, and S.P.A. Bordas. A tutorial on bayesian inference to identify material parameters in solid mechanics. Archives of Computational Methods in Engineering, 27(2):361–385, 2020.
Gishan D. Ranasinghe, Tony Lindgren, Mark Girolami, and Ajith K. Parlikad. A methodology for prognostics under the conditions of limited failure data availability. IEEE Access, 7:183996–184007, 2019. URL: https://doi.org/10.1109%2Faccess.2019.2960310, doi:10.1109/access.2019.2960310.
C. Ye, L. Butler, B. Calka, M. Iangurazov, Q. Lu, A. Gregory, M. Girolami, and C. Middleton. A digital twin of bridges for structural health monitoring. Structural Health Monitoring 2019: Enabling Intelligent Life-Cycle Health Management for Industry Internet of Things (IIOT) - Proceedings of the 12th International Workshop on Structural Health Monitoring, 1:1619–1626, 2019.
A. Mendoza, L. Roininen, M. Girolami, J. Heikkinen, and H. Haario. Accelerated whole-core analysis optimization with wellsite tomography instrumentation and bayesian inversion. Petrophysics, 60(3):384–396, 2019.
C.J. Oates, J. Cockayne, R.G. Aykroyd, and M. Girolami. Bayesian probabilistic numerical methods in time-dependent state estimation for industrial hydrocyclone equipment. Journal of the American Statistical Association, 114(528):1518–1531, 2019.
J. Cockayne, C.J. Oates, T.J. Sullivan, and M. Girolami. Bayesian probabilistic numerical methods. SIAM Review, 61(4):756–789, 2019.
C.J. Oates, J. Cockayne, F.-X. Briol, and M. Girolami. Convergence rates for a class of estimators based on stein?s method. Bernoulli, 25(2):1141–1159, 2019.
M. Girolami, I.C.F. Ipsen, C.J. Oates, A.B. Owen, and T.J. Sullivan. Editorial: special edition on probabilistic numerics. Statistics and Computing, 29(6):1181–1183, 2019.
R. Scalzo, D. Kohn, H. Olierook, G. Houseman, R. Chandra, M. Girolami, and S. Cripps. Efficiency and robustness in monte carlo sampling for 3-d geophysical inversions with obsidian v0.1.2: setting up for success. Geoscientific Model Development, 12(7):2941–2960, 2019.
L. Roininen, M. Girolami, S. Lasanen, and M. Markkanen. Hyperpriors for matérn fields with applications in bayesian inversion. Inverse Problems and Imaging, 13(1):1–29, 2019.
A. Barp, F.-X. Briol, A.B. Duncan, M. Girolami, and L. Mackey. Minimum stein discrepancy estimators. Advances in Neural Information Processing Systems, 2019.
O. Hamelijnck, T. Damoulas, K. Wang, and M.A. Girolami. Multi-resolution multi-task gaussian processes. Advances in Neural Information Processing Systems, 2019.
S. Livingstone, M. Betancourt, S. Byrne, and M. Girolami. On the geometric ergodicity of hamiltonian monte carlo. Bernoulli, 25(4 A):3109–3138, 2019.
S. Virtanen and M. Girolami. Precision-recall balanced topic modelling. Advances in Neural Information Processing Systems, 2019.
F.-X. Briol, C.J. Oates, M. Girolami, M.A. Osborne, and D. Sejdinovic. Probabilistic integration: a role in statistical computation? Statistical Science, 34(1):1–22, 2019.
F.-X. Briol, C.J. Oates, M. Girolami, M.A. Osborne, and D. Sejdinovic. Rejoinder: probabilistic integration: a role in statistical computation? Statistical Science, 34(1):38–42, 2019.
A. Mendoza, L. Roininen, M. Girolami, J. Heikkinen, and H. Haario. Statistical methods to enable practical on-site tomographic imaging of whole-core samples. Geophysics, 84(3):D89–D100, 2019.
W.Y. Chen, A. Barp, F.-X. Briol, J. Gorham, M. Girolami, L. Mackey, and C.J. Oates. Stein point markov chain monte carlo. 36th International Conference on Machine Learning, ICML 2019, 2019-June:1737–1767, 2019.
A. Gregory, F.D.-H. Lau, M. Girolami, L.J. Butler, and M.Z.E.B. Elshafie. The synthesis of data from instrumented structures and physics-based models via gaussian processes. Journal of Computational Physics, 392:248–265, 2019.
A. Manderson, M.D. Rayson, E. Cripps, M. Girolami, J.P. Gosling, M. Hodkiewicz, G.N. Ivey, and N.L. Jones. Uncertainty quantification of density and stratification estimates with implications for predicting ocean dynamics. Journal of Atmospheric and Oceanic Technology, 36(8):1313–1330, 2019.
L.A. Bull, T.J. Rogers, C. Wickramarachchi, E.J. Cross, K. Worden, and N. Dervilis. Probabilistic active learning: an online framework for structural health monitoring. Mechanical Systems and Signal Processing, 134:106294, dec 2019. URL: https://doi.org/10.1016%2Fj.ymssp.2019.106294, doi:10.1016/j.ymssp.2019.106294.
L.A. Bull, K. Worden, R. Fuentes, G. Manson, E.J. Cross, and N. Dervilis. Outlier ensembles: a robust method for damage detection and unsupervised feature extraction from high-dimensional data. Journal of Sound and Vibration, 453:126–150, aug 2019. URL: https://doi.org/10.1016%2Fj.jsv.2019.03.025, doi:10.1016/j.jsv.2019.03.025.
LA Bull, K Worden, TJ Rogers, C Wickramarachchi, EJ Cross, T McLeay, W Leahy, and N Dervilis. A probabilistic framework for online structural health monitoring : active learning from machining data streams. In Journal of Physics: Conference Series, volume 1264. Valpre, Lyon, France, IOP Publishing, Jul 2019. doi:10.1088/1742-6596/1264/1/012028.
N Dervilis, T Zhang, L Bull, EJ Cross, TJ Rogers, R Fuentes, V Dertimanis, I Abdallah, E Chatzi, and K Worden. A nonlinear robust outlier detection approach for shm. In 8th IOMAC - International Operational Modal Analysis Conference, Proceedings, 107–114. Jan 2019.
L Bull, G Manson, K Worden, and N Dervilis. Active learning approaches to structural health monitoring. In 157–159. Springer International Publishing, 2019. doi:10.1007/978-3-319-75390-4_14.
Toni Karvonen and Simo Särkkä. Gaussian kernel quadrature at scaled gauss–hermite nodes. BIT Numerical Mathematics, 59(4):877–902, jun
Filip Tronarp, Toni Karvonen, and Simo Sarkka. Students $\less$inline-formula$\greater$ $\less$tex-math notation="LaTeX"$\greater$t$\less$/tex-math$\greater$ $\less$/inline-formula$\greater$-filters for noise scale estimation. IEEE Signal Processing Letters, 26(2):352–356, feb 2019. URL: https://doi.org/10.1109%2Flsp.2018.2889440, doi:10.1109/lsp.2018.2889440.
Ieva Kazlauskaite, Carl Henrik Ek, and Neill DF Campbell. Gaussian process latent variable alignment learning. In International Conference on Artificial Intelligence and Statistics (AISTATS 2019). Proceedings of Machine Learning Research, 2019.
Erik Bodin, Markus Kaiser, Ieva Kazlauskaite, Zhenwen Dai, Neill DF Campbell, and Carl Henrik Ek. Modulating surrogates for bayesian optimization. In International Conference on Machine Learning (ICML 2020), arXiv–1906. 2019.
H. Rappel, L. Wu, L. Noels, and L.A.A. Beex. A bayesian framework to identify random parameter fields based on the copula theorem and gaussian fields: application to polycrystalline materials. Journal of Applied Mechanics, Transactions ASME, 2019.
M. Mohamedou, K. Zulueta, C.N. Chung, H. Rappel, L. Beex, L. Adam, A. Arriaga, Z. Major, L. Wu, and L. Noels. Bayesian identification of mean-field homogenization model parameters and uncertain matrix behavior in non-aligned short fiber composites. Composite Structures, 220:64–80, 2019.
H. Rappel and L.A.A. Beex. Estimating fibres? material parameter distributions from limited data with the help of bayesian inference. European Journal of Mechanics, A/Solids, 75:169–196, 2019.
H. Rappel, L.A.A. Beex, L. Noels, and S.P.A. Bordas. Identifying elastoplastic parameters with bayes? theorem considering output error, input error and model uncertainty. Probabilistic Engineering Mechanics, 55:28–41, 2019.
O. Mac Aodha, R. Gibb, K.E. Barlow, E. Browning, M. Firman, R. Freeman, B. Harder, L. Kinsey, G.R. Mead, S.E. Newson, I. Pandourski, S. Parsons, J. Russ, A. Szodoray-Paradi, F. Szodoray-Paradi, E. Tilova, M. Girolami, G. Brostow, and K.E. Jones. Bat detective?deep learning tools for bat acoustic signal detection. PLoS Computational Biology, 2018.
V. Stathopoulos, V. Zamora-Gutierrez, K.E. Jones, and M. Girolami. Bat echolocation call identification for biodiversity monitoring: a probabilistic approach. Journal of the Royal Statistical Society. Series C: Applied Statistics, 67(1):165–183, 2018.
X. Xi, F.-X. Briol, and M. Girolami. Bayesian quadrature for multiple related integrals. 35th International Conference on Machine Learning, ICML 2018, 12:8533–8564, 2018.
A. Barp, F.-X. Briol, A.D. Kennedy, and M. Girolami. Geometry and dynamics for markov chain monte carlo. Annual Review of Statistics and Its Application, 5:451–471, 2018.
M.M. Dunlop, M.A. Girolami, A.M. Stuart, and A.L. Teckentrup. How deep are deep gaussian processes? Journal of Machine Learning Research, 19:1–46, 2018.
J. Noonan, S.M. Asiala, G. Grassia, N. MacRitchie, K. Gracie, J. Carson, M. Moores, M. Girolami, A.C. Bradshaw, T.J. Guzik, G.R. Meehan, H.E. Scales, J.M. Brewer, I.B. McInnes, N. Sattar, K. Faulds, P. Garside, D. Graham, and P. Maffia. In vivo multiplex molecular imaging of vascular inflammation using surface-enhanced raman spectroscopy. Theranostics, 8(22):6195–6209, 2018.
A. Mendoza, L. Roininen, M. Girolami, J. Heikkinen, and H. Haario. Statistical methods to enable practical on-site tomographic imaging of whole-core samples. SPWLA 59th Annual Logging Symposium 2018, 2018.
L. Ellam, M. Girolami, G.A. Pavliotis, and A. Wilson. Stochastic modelling of urban structure. Proceedings of the Royal Society A: Mathematical, Physical and Engineering Sciences, 2018.
F.D.-H. Lau, N.M. Adams, M.A. Girolami, L.J. Butler, and M.Z.E.B. Elshafie. The role of statistics in data-centric engineering. Statistics and Probability Letters, 136:58–62, 2018.
L. Bull, K. Worden, G. Manson, and N. Dervilis. Active learning for semi-supervised structural health monitoring. Journal of Sound and Vibration, 437:373–388, dec 2018. URL: https://doi.org/10.1016%2Fj.jsv.2018.08.040, doi:10.1016/j.jsv.2018.08.040.
H. Rappel, L.A.A. Beex, and S.P.A. Bordas. Bayesian inference to identify parameters in viscoelasticity. Mechanics of Time-Dependent Materials, 22(2):221–258, 2018.
L. Ellam, H. Strathmann, M. Girolami, and I. Murray. A determinant-free method to simulate the parameters of large gaussian fields. Stat, 6(1):271–281, 2017.
K. Jensen, C. Soguero-Ruiz, K. Oyvind Mikalsen, R.-O. Lindsetmo, I. Kouskoumvekaki, M. Girolami, S. Olav Skrovseth, and K. Magne Augestad. Analysis of free text in electronic health records for identification of cancer patient trajectories. Scientific Reports, 2017.
C.J. Oates, M. Girolami, and N. Chopin. Control functionals for monte carlo integration. Journal of the Royal Statistical Society. Series B: Statistical Methodology, 79(3):695–718, 2017.
A. Beskos, M. Girolami, S. Lan, P.E. Farrell, and A.M. Stuart. Geometric mcmc for infinite-dimensional inverse problems. Journal of Computational Physics, 335:327–351, 2017.
F.-X. Briol, C.J. Oates, J. Cockayne, W.Y. Chen, and M. Girolami. On the sampling problem for kernel quadrature. 34th International Conference on Machine Learning, ICML 2017, 2:949–968, 2017.
C.J. Oates, S. Niederer, A. Lee, F.-X. Briol, and M. Girolami. Probabilistic models for integration error in the assessment of functional cardiac models. Advances in Neural Information Processing Systems, 2017-December:110–118, 2017.
J. Cockayne, C. Oates, T. Sullivan, and M. Girolami. Probabilistic numerical methods for pde-constrained bayesian inverse problems. AIP Conference Proceedings, 2017.
P.R. Conrad, M. Girolami, S. Särkkä, A. Stuart, and K. Zygalakis. Statistical analysis of differential equations: introducing probability measures on numerical solutions. Statistics and Computing, 27(4):1065–1082, 2017.
M. Betancourt, S. Byrne, S. Livingstone, and M. Girolami. The geometric foundations of hamiltonian monte carlo. Bernoulli, 23(4A):2257–2298, 2017.
L. Ellam, N. Zabaras, and M. Girolami. A bayesian approach to multiscale inverse problems with on-the-fly scale determination. Journal of Computational Physics, 326:115–140, 2016.
M. Epstein, B. Calderhead, M.A. Girolami, and L.G. Sivilotti. Bayesian statistical inference in ion-channel models with exact missed event correction. Biophysical Journal, 111(2):333–348, 2016.
O.A. Chkrebtii, D.A. Campbell, B. Calderhead, and M.A. Girolami. Bayesian solution uncertainty quantification for differential equations. Bayesian Analysis, 11(4):1239–1267, 2016.
T. House, A. Ford, S. Lan, S. Bilson, E. Buckingham-Jeffery, and M. Girolami. Bayesian uncertainty quantification for transmissibility of influenza, norovirus and ebola using information geometry. Journal of the Royal Society Interface, 2016.
C.J. Oates and M. Girolami. Control functionals for quasi-monte carlo integration. Proceedings of the 19th International Conference on Artificial Intelligence and Statistics, AISTATS 2016, pages 56–65, 2016.
J.M. Rondina, M. Filippone, M. Girolami, and N.S. Ward. Decoding post-stroke motor function from structural brain imaging. NeuroImage: Clinical, 12:372–380, 2016.
S. Lan, T. Bui-Thanh, M. Christie, and M. Girolami. Emulation of higher-order tensors in manifold monte carlo methods for bayesian inverse problems. Journal of Computational Physics, 308:81–101, 2016.
K. Gracie, M. Moores, W.E. Smith, K. Harding, M. Girolami, D. Graham, and K. Faulds. Preferential attachment of specific fluorescent dyes and dye labeled dna sequences in a surface enhanced raman scattering multiplex. Analytical Chemistry, 88(2):1147–1153, 2016.
B. Banushi, F. Forneris, A. Straatman-Iwanowska, A. Strange, A.-M. Lyne, C. Rogerson, J.J. Burden, W.E. Heywood, J. Hanley, I. Doykov, K.R. Straatman, H. Smith, D. Bem, J. Kriston-Vizi, G. Ariceta, M. Risteli, C. Wang, R.E. Ardill, M. Zaniew, J. Latka-Grot, S.N. Waddington, S.J. Howe, F. Ferraro, A. Gjinovci, S. Lawrence, M. Marsh, M. Girolami, L. Bozec, K. Mills, and P. Gissen. Regulation of post-golgi lh3 trafficking is essential for collagen homeostasis. Nature Communications, 2016.
P.S. Koutsourelakis, N. Zabaras, and M. Girolami. Special issue: big data and predictive computational modeling. Journal of Computational Physics, 321:1252–1254, 2016.
C.J. Oates, T. Papamarkou, and M. Girolami. The controlled thermodynamic integral for bayesian model evidence evaluation. Journal of the American Statistical Association, 111(514):634–645, 2016.
S. Virtanen, M. Rost, A. Morrison, M. Chalmers, and M. Girolami. Uncovering smartphone usage patterns with multi-view mixed membership models. Stat, 5(1):57–69, 2016.
Toni Karvonen and Simo Sarkka. Approximate state-space gaussian processes via spectral transformation. In 2016 IEEE 26th International Workshop on Machine Learning for Signal Processing (MLSP). Institute of Electrical and Electronics Engineers (IEEE), sep 2016. URL: https://doi.org/10.1109%2Fmlsp.2016.7738812, doi:10.1109/mlsp.2016.7738812.
L.E.M. Hopcroft, B. Calderhead, P. Gallipoli, T.L. Holyoake, and M.A. Girolami. Bayesian inference for model selection: an application to aberrant signalling pathways in chronic myeloid leukaemia. Systems Genetics: Linking Genotypes and Phenotypes, pages 161–190, 2015.
R. Schwentner, T. Papamarkou, M.O. Kauer, V. Stathopoulos, F. Yang, S. Bilke, P.S. Meltzer, M. Girolami, and H. Kovar. Ews-fli1 employs an e2f switch to drive target gene expression. Nucleic Acids Research, 43(5):2780–2789, 2015.
F.-X. Briol, C.J. Oates, M. Girolami, and M.A. Osborne. Frank-wolfe bayesian quadrature: probabilistic integration with theoretical guarantees. Advances in Neural Information Processing Systems, 2015-January:1162–1170, 2015.
S. Lan, V. Stathopoulos, B. Shahbaba, and M. Girolami. Markov chain monte carlo from lagrangian dynamics. Journal of Computational and Graphical Statistics, 24(2):357–378, 2015.
S. Virtanen, M. Rost, M. Higgs, A. Morrison, M. Chalmers, and M. Girolami. Non-parametric bayes to infer playing strategies adopted in a population of mobile gamers. Stat, 4(1):46–58, 2015.
A.-M. Lyne, M. Girolami, Y. Atchadé, H. Strathmann, and D. Simpson. On russian roulette estimates for bayesian inference with doubly-intractable likelihoods. Statistical Science, 30(4):443–467, 2015.
S. Virtanen and M. Girolami. Ordinal mixed membership models. 32nd International Conference on Machine Learning, ICML 2015, 1:588–596, 2015.
P. Hennig, M.A. Osborne, and M. Girolami. Probabilistic numerics and uncertainty in computations. Proceedings of the Royal Society A: Mathematical, Physical and Engineering Sciences, 2015.
Alessandro Barp, Edoardo Gabriele Barp, François-Xavier Briol, and Daniel Ueltschi. A numerical study of the 3d random interchange and random loop models. Journal of Physics A: Mathematical and Theoretical, 48(34):345002, aug 2015. URL: https://doi.org/10.1088%2F1751-8113%2F48%2F34%2F345002, doi:10.1088/1751-8113/48/34/345002.
V. Stathopoulos, V. Zamora-Gutierrez, K.E. Jones, and M. Girolami. Bat call identification with gaussian process multinomial probit regression and a dynamic time warping kernel. Journal of Machine Learning Research, 33:913–921, 2014.
M.A. Girolami. Contribution by m. a. girolami. Statistical Science, 29(1):97, 2014.
S. Livingstone and M. Girolami. Information-geometric markov chain monte carlo methods using diffusions. Entropy, 16(6):3074–3102, 2014.
T. Xifara, C. Sherlock, S. Livingstone, S. Byrne, and M. Girolami. Langevin diffusions and the metropolis-adjusted langevin algorithm. Statistics and Probability Letters, 91(1):14–19, 2014.
A. Kramer, V. Stathopoulos, M. Girolami, and N. Radde. Mcmc_clib-an advanced mcmc sampling package for ode models. Bioinformatics (Oxford, England), 30(20):2991–2992, 2014.
O. Andrei, M. Calder, M. Higgs, and M. Girolami. Probabilistic model checking of dtmc models of user activity patterns. Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), 8657 LNCS:138–153, 2014.
M. Filippone and M. Girolami. Pseudo-marginal bayesian inference for gaussian processes. IEEE Transactions on Pattern Analysis and Machine Intelligence, 36(11):2214–2226, 2014.
O.M. Aodha, V. Stathopoulos, G.J. Brostow, M. Terry, M. Girolami, and K.E. Jones. Putting the scientist in the loop - accelerating scientific progress with interactive machine learning. Proceedings - International Conference on Pattern Recognition, pages 9–17, 2014.
M. Jiwaji, M.E. Sandison, J. Reboud, R. Stevenson, R. Daly, G. Barkess, K. Faulds, W. Kolch, D. Graham, M.A. Girolami, J.M. Cooper, and A.R. Pitt. Quantification of functionalised gold nanoparticle-targeted knockdown of gene expression in hela cells. PLoS ONE, 2014.
S. Byrne and M. Girolami. Rejoinder: geodesic monte carlo on embedded manifolds. Scandinavian Journal of Statistics, 41(1):19–21, 2014.
T. Bui-Thanh and M. Girolami. Solving large-scale pde-constrained bayesian inverse problems with riemann manifold hamiltonian monte carlo. Inverse Problems, 2014.
F. Achcar, A. Fadda, J.R. Haanstra, E.J. Kerkhoven, D.-H. Kim, A.E. Leroux, T. Papamarkou, F. Rojas, B.M. Bakker, M.P. Barrett, C. Clayton, M. Girolami, R.L. Krauth-Siegel, K.R. Matthews, and R. Breitling. The silicon trypanosome. a test case of iterative model extension in systems biology. Advances in Microbial Physiology, 64:115–143, 2014.
T. Papamarkou, A. Mira, and M. Girolami. Zero variance differential geometric markov chain monte carlo algorithms. Bayesian Analysis, 9(1):97–128, 2014.
H. Rappel, A. Yousefi-Koma, J. Jamali, and A. Bahari. Numerical time-domain modeling of lamb wave propagation using elastodynamic finite integration technique. Shock and Vibration, 2014.
M. Filippone, M. Zhong, and M. Girolami. A comparative evaluation of stochastic-based inference methods for gaussian process models. Machine Learning, 93(1):93–114, 2013.
M. Higgs, A. Morrison, M. Girolami, and M. Chalmers. Analysing user behaviour through dynamic population models. Conference on Human Factors in Computing Systems - Proceedings, 2013-April:271–276, 2013.
A.F. Marquand, M. Filippone, J. Ashburner, M. Girolami, J. Mourao-Miranda, G.J. Barker, S.C.R. Williams, P.N. Leigh, and C.R.V. Blain. Automated, high accuracy classification of parkinsonian disorders: a pattern recognition approach. PLoS ONE, 2013.
B. Calderhead, M. Epstein, L. Sivilotti, and M. Girolami. Bayesian approaches for mechanistic ion channel modeling. Methods in Molecular Biology, 1021:247–272, 2013.
M. Girolami and A. Mira. Comment on article by schmidl et al. Bayesian Analysis, 8(1):27–32, 2013.
S. Byrne and M. Girolami. Geodesic monte carlo on embedded manifolds. Scandinavian Journal of Statistics, 40(4):825–845, 2013.
V. Stathopoulos and M.A. Girolami. Markov chain monte carlo inference for markov jump processes via the linear noise approximation. Philosophical Transactions of the Royal Society A: Mathematical, Physical and Engineering Sciences, 2013.
T. Diethe and M. Girolami. Online learning with (multiple) kernels: a review. Neural Computation, 25(3):567–625, 2013.
M. Zhong and M. Girolami. A bayesian approach to approximate joint diagonalization of square matrices. Proceedings of the 29th International Conference on Machine Learning, ICML 2012, 1:647–654, 2012.
K. Yuan, M. Girolami, and M. Niranjan. Markov chain monte carlo methods for state-space models with point process observations. Neural Computation, 24(6):1462–1486, 2012.
L. Mohamed, B. Calderhead, M. Filippone, M. Christie, and M. Girolami. Population mcmc methods for history matching and uncertainty quantification. Computational Geosciences, 16(2):423–436, 2012.
N. Lawrence and M. Girolami. Preface. Journal of Machine Learning Research, 2012.
M. Filippone, A.F. Marquand, C.R.V. Blain, S.C.R. Williams, J. Mourão-Miranda, and M. Girolami. Probabilistic prediction of neurological disorders with a statistical assessment of neuroimaging data modalities. Annals of Applied Statistics, 6(4):1883–1905, 2012.
M. Jiwaji, R. Daly, A. Gibriel, G. Barkess, P. McLean, J. Yang, K. Pansare, S. Cumming, A. McLauchlan, P.J. Kamola, M.S. Bhutta, A.G. West, K.L. West, W. Kolch, M.A. Girolami, and A.R. Pitt. Unique reporter-based sensor platforms to monitor signalling in cells. PLoS ONE, 2012.
S. Rogers and M. Girolami. A first course in machine learning. A First Course in Machine Learning, pages 1–284, 2011.
M. Zhong, M. Girolami, K. Faulds, and D. Graham. Bayesian methods to detect dye-labelled dna oligonucleotides in multiplexed raman spectra. Journal of the Royal Statistical Society. Series C: Applied Statistics, 60(2):187–206, 2011.
M. Filippone, A. Mira, and M. Girolami. Discussion of the paper: "sampling schemes for generalized linear dirichlet process random effects models" by m. kyung, j. gill, and g. casella. Statistical Methods and Applications, 20(3):295–297, 2011.
M.P.H. Stumpf, D.J. Balding, and M. Girolami. Handbook of statistical systems biology. Handbook of Statistical Systems Biology, pages 1–508, 2011.
H. Lehrach, R. Subrak, P. Boyle, M. Pasterk, K. Zatloukal, H. Müller, T. Hubbard, A. Brand, M. Girolami, D. Jameson, F.J. Bruggeman, and H.V. Westerhoff. Itfom - the it future of medicine. Procedia Computer Science, 7:26–29, 2011.
T. Honkela, W. Duch, M. Girolami, and S. Kaski. Lecture notes in computer science (including subseries lecture notes in artificial intelligence and lecture notes in bioinformatics): preface. Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), 2011.
V. Stathopoulos and M. Girolami. Manifold mcmc for mixtures. Mixtures: Estimation and Applications, pages 255–276, 2011.
R. Paredes and M. Girolami. On the use of diagonal and class-dependent weighted distances for the probabilistic k-nearest neighbor. Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), 6669 LNCS:265–272, 2011.
T. Polajnar, S. Rogers, and M. Girolami. Protein interaction detection in sentences via gaussian processes: a preliminary evaluation. International Journal of Data Mining and Bioinformatics, 5(1):52–72, 2011.
T. Polajnar, T. Damoulas, and M. Girolami. Protein interaction sentence detection using multiple semantic kernels. Journal of Biomedical Semantics, 2011.
M. Girolami and B. Calderhead. Riemann manifold langevin and hamiltonian monte carlo methods. Journal of the Royal Statistical Society. Series B: Statistical Methodology, 73(2):123–214, 2011.
B. Calderhead and M. Girolami. Statistical analysis of nonlinear dynamical systems using differential geometric sampling methods. Interface Focus, 1(6):821–835, 2011.
M. Dakna, K. Harris, A. Kalousis, S. Carpentier, W. Kolch, J.P. Schanstra, M. Haubitz, A. Vlahou, H. Mischak, and M. Girolami. Addressing the challenge of defining valid proteomic biomarkers and classifiers. BMC Bioinformatics, 2010.
T.-R. Xu, V. Vyshemirsky, A. Gormand, A. Von Kriegsheim, M. Girolami, G.S. Baillie, D. Ketley, A.J. Dunlop, G. Milligan, M.D. Houslay, and W. Kolch. Inferring signaling pathway topologies from multiple perturbation measurements of specific biochemical species. Science Signaling, 2010.
T.-R. Xu, V. Vyshemirsky, A. Gormand, A. Von Kriegsheim, M. Girolami, G.S. Baillie, D. Ketley, A.J. Dunlop, G. Milligan, M.D. Houslay, and W. Kolch. Inferring signaling pathway topologies from multiple perturbation measurements of specific biochemical species (science signaling (2010) 3, (er8)). Science Signaling, 2010.
S. Rogers, A. Klami, J. Sinkkonen, M. Girolami, and S. Kaski. Infinite factorization of multiple non-parametric views. Machine Learning, 79(1-2):201–226, 2010.
I. Psorakis, T. Damoulas, and M.A. Girolami. Multiclass relevance vector machines: sparsity and accuracy. IEEE Transactions on Neural Networks, 21(10):1588–1598, 2010.
D.M. Good, P. Zürbig, ?. Argilés, H.W. Bauer, G. Behrens, J.J. Coon, M. Dakna, S. Decramer, C. Delles, A.F. Dominiczak, J.H.H. Ehrich, F. Eitner, D. Fliser, M. Frommberger, A. Ganser, M.A. Girolami, I. Golovko, W. Gwinner, M. Haubitz, S. Herget-Rosenthal, J. Jankowski, H. Jahn, G. Jerums, B.A. Julian, M. Kellmann, V. Kliem, W. Kolch, A.S. Krolewski, M. Luppi, Z. Massy, M. Melter, C. Neusüss, J. Novak, K. Peter, K. Rossing, H. Rupprecht, J.P. Schanstra, E. Schiffer, J.-U. Stolzenburg, L. Tarnow, D. Theodorescu, V. Thongboonkerd, R. Vanholder, E.M. Weissinger, H. Mischak, and P. Schmitt-Kopplinr. Naturally occurring human urinary peptides for use in diagnosis of chronic kidney disease. Molecular and Cellular Proteomics, 9(11):2424–2437, 2010.
L. Mohamed, B. Calderhead, M. Filippone, M. Christie, and M. Girolami. Population mcmc methods for history matching and uncertainty quantification. ECMOR 2010 - 12th European Conference on the Mathematics of Oil Recovery, 2010.
L.E.M. Hopcroft, M.W. McBride, K.J. Harris, A.K. Sampson, J.D. McClure, D. Graham, G. Young, T.L. Holyoake, M.A. Girolami, and A.F. Dominiczak. Predictive response-relevant clustering of expression data provides insights into disease processes. Nucleic Acids Research, 38(20):6831–6840, 2010.
H. Mischak, G. Allmaier, R. Apweiler, T. Attwood, M. Baumann, A. Benigni, S.E. Bennett, R. Bischoff, E. Bongcam-Rudloff, G. Capasso, J.J. Coon, P. D’Haese, A.F. Dominiczak, M. Dakna, H. Dihazi, J.H. Ehrich, P. Fernandez-Llama, D. Fliser, J. Frokiaer, J. Garin, M. Girolami, W.S. Hancock, M. Haubitz, D. Hochstrasser, R.R. Holman, J.P.A. Ioannidis, J. Jankowski, B.A. Julian, J.B. Klein, W. Kolch, T. Luider, Z. Massy, W.B. Mattes, F. Molina, B. Monsarrat, J. Novak, K. Peter, P. Rossing, M. Sánchez-Carbayo, J.P. Schanstra, O.J. Semmes, G. Spasovski, D. Theodorescu, V. Thongboonkerd, R. Vanholder, T.D. Veenstra, E. Weissinger, T. Yamamoto, and A. Vlahou. Recommendations for biomarker identification and qualification in clinical proteomics. Science Translational Medicine, 2010.
S. Rogers, M. Girolami, and T. Polajnar. Semi-parametric analysis of multi-rater data. Statistics and Computing, 20(3):317–334, 2010.
A.F. Dominiczak, S. Herget-Rosenthal, C. Delles, D. Fliser, I. Fournier, A. Graber, M. Girolami, E. Holmes, F. Lang, F. Molina, J. Nicholson, G. Remuzzi, P. Rossing, K.L. Rudolph, O. Wolkenhauer, I. Xenarios, R. Zubarev, D. Zubov, A. Vlahou, and J.P. Schanstra. Systems biology to battle vascular disease. Nephrology Dialysis Transplantation, 25(4):1019–1022, 2010.
F. Molina, M. Dehmer, P. Perco, A. Graber, M. Girolami, G. Spasovski, J.P. Schanstra, and A. Vlahou. Systems biology: opening new avenues in clinical research. Nephrology Dialysis Transplantation, 25(4):1015–1018, 2010.
B.M. Bakker, R.L. Krauth-Siegel, C. Clayton, K. Matthews, M. Girolami, H.V. Westerhoff, P.A.M. Michels, R. Breitling, and M.P. Barrett. The silicon trypanosome. Parasitology, 137(9):1333–1341, 2010.
B. Calderhead, M. Girolami, and N.D. Lawrence. Accelerating bayesian inference over nonlinear differential equations with gaussian processes. Advances in Neural Information Processing Systems 21 - Proceedings of the 2008 Conference, pages 217–224, 2009.
Y. Ying, C. Campbell, and M. Girolami. Analysis of svm with indefinite kernels. Advances in Neural Information Processing Systems 22 - Proceedings of the 2009 Conference, pages 2205–2213, 2009.
Y. Ying, C. Campbell, T. Damoulas, and M. Girolami. Class prediction from disparate biological data sources using an iterative multi-kernel algorithm. Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), 5780 LNBI:427–438, 2009.
T. Polajnar, S. Rogers, and M. Girolami. Classification of protein interaction sentences via gaussian processes. Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), 5780 LNBI:282–292, 2009.
T. Damoulas and M.A. Girolami. Combining feature spaces for classification. Pattern Recognition, 42(11):2671–2683, 2009.
K. Harris, M. Girolami, and H. Mischak. Definition of valid proteomic biomarkers: a bayesian solution. Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), 5780 LNBI:137–149, 2009.
B. Calderhead and M. Girolami. Estimating bayes factors via thermodynamic integration and population mcmc. Computational Statistics and Data Analysis, 53(12):4028–4045, 2009.
K. Harris, L. McMillan, and M. Girolami. Inferring meta-covariates in classification. Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), 5780 LNBI:150–161, 2009.
M. Girolami, V. Kadirkamanathan, M. Niranjan, J. Noirel, and G. Sanguinetti. Lecture notes in computer science (including subseries lecture notes in artificial intelligence and lecture notes in bioinformatics): preface. Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), 2009.
T. Damoulas and M.A. Girolami. Pattern recognition with a bayesian kernel combination machine. Pattern Recognition Letters, 30(1):46–54, 2009.
S. Rogers, R.A. Scheltema, M. Girolami, and R. Breitling. Probabilistic assignment of formulas to mass peaks in metabolomics experiments. Bioinformatics, 25(4):512–518, 2009.
M. Girolami. Reversible jump mcmc for non-negative matrix factorization with application to raman spectral decomposition. Proceedings of SPIE - The International Society for Optical Engineering, 2009.
M. Zhong and M. Girolami. Reversible jump mcmc for non-negative matrix factorization. Journal of Machine Learning Research, 5:663–670, 2009.
T. Polajnar and M. Girolami. Semi-supervised prediction of protein interaction sentences exploiting semantically encoded metrics. Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), 5780 LNBI:270–281, 2009.
R. Daly, K.D. Edwards, J.S. O’Neill, S. Aitken, A.J. Millar, and M. Girolami. Using higher-order dynamic bayesian networks to model periodic data from the circadian clock of arabidopsis thaliana. Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), 5780 LNBI:67–78, 2009.
N. Waddell, A. Ten Haaf, A. Marsh, J. Johnson, L.C. Walker, M. Gongora, M. Brown, P. Grover, M. Girolami, S. Grimmond, G. Chenevix-Trench, and A.B. Spurdle. Brca1 and brca2 missense variants of high and low clinical significance influence lymphoblastoid cell line post-irradiation gene expression. PLoS Genetics, 2008.
M. Girolami. Bayesian inference for differential equations. Theoretical Computer Science, 408(1):4–16, 2008.
V. Vyshemirsky and M.A. Girolami. Bayesian ranking of biochemical system models. Bioinformatics, 24(6):833–839, 2008.
V. Vyshemirsky and M. Girolami. Biobayes: a software package for bayesian inference in systems biology. Bioinformatics, 24(17):1933–1934, 2008.
M. Zhong, F. Lotte, M. Girolami, and A. Lécuyer. Classifying eeg for brain computer interfaces using gaussian processes. Pattern Recognition Letters, 29(3):354–359, 2008.
V. Vyshemirsky and M.A. Girolami. Erratum: bayesian ranking of biochemical system models (bioinformatics (2008) vol. 24(6) (833-839)). Bioinformatics, 24(20):2421, 2008.
T. Damoulas, Y. Ying, M.A. Girolami, and C. Campbell. Inferring sparse kernel combinations and relevance vectors: an application to subcellular localization of proteins. Proceedings - 7th International Conference on Machine Learning and Applications, ICMLA 2008, pages 577–582, 2008.
S. Rogers, M. Girolami, W. Kolch, K.M. Waters, T. Liu, B. Thrall, and S.H. Wiley. Investigating the correspondence between transcriptomic and proteomic expression profiles using coupled cluster models. Bioinformatics, 24(24):2894–2900, 2008.
I.M. Overton, G. Padovani, M.A. Girolami, and G.J. Barton. Parcrys: a parzen window density estimation approach to protein crystallization propensity prediction. Bioinformatics, 24(7):901–907, 2008.
T. Damoulas and M.A. Girolami. Probabilistic multi-class multi-kernel learning: on protein fold recognition and remote homology detection. Bioinformatics, 24(10):1264–1270, 2008.
N. Lama and M. Girolami. Vbmp: variational bayesian multinomial probit regression for multi-class classification in r. Bioinformatics, 24(1):135–136, 2008.
D. Fliser, J. Novak, V. Thongboonkerd, A. Argiles, V. Jankowski, M.A. Girolami, J. Jankowski, and H. Mischak. Advances in urinary proteome analysis and biomarker discovery. Journal of the American Society of Nephrology, 18(4):1057–1071, 2007.
S. Manocha and M.A. Girolami. An empirical analysis of the probabilistic k-nearest neighbour classifier. Pattern Recognition Letters, 28(13):1818–1824, 2007.
S. Rogers, R. Khanin, and M. Girolami. Bayesian model-based inference of transcription factor activity. BMC Bioinformatics, 2007.
H. Mischak, R. Apweiler, R.E. Banks, M. Conaway, J. Coon, A. Dominiczak, J.H.H. Ehrich, D. Fliser, M. Girolami, H. Hermjakob, D. Hochstrasser, J. Jankowski, B.A. Julian, W. Kolch, Z.A. Massy, C. Neusuess, J. Novak, K. Peter, K. Rossing, J. Schanstra, O.J. Semmes, D. Theodorescu, V. Thongboonkerd, E.M. Weissinger, J.E. Van Eyk, and T. Yamamoto. Clinical proteomics: a need to define the field and to begin to set adequate standards. Proteomics - Clinical Applications, 1(2):148–156, 2007.
M. Girolami and M. Zhong. Data integration for classification problems employing gaussian process priors. Advances in Neural Information Processing Systems, pages 465–472, 2007.
O. Sharma, M. Girolami, and J. Sventek. Detecting worm variants using machine learning. Proceedings of 2007 ACM CoNEXT Conference - 3rd International Conference on Emerging Networking EXperiments and Technologies, CoNEXT, 2007.
D. Xing and M. Girolami. Employing latent dirichlet allocation for fraud detection in telecommunications. Pattern Recognition Letters, 28(13):1727–1734, 2007.
R. Jenssen, T. Eltoft, M. Girolami, and D. Erdogmus. Kernel maximum entropy data transformation and an enhanced spectral clustering algorithm. Advances in Neural Information Processing Systems, pages 633–640, 2007.
G.C. Cawley, N.L.C. Talbot, and M. Girolami. Sparse multinomial logistic regression via bayesian l1 regularisation. Advances in Neural Information Processing Systems, pages 209–216, 2007.
Z. Kote-Jarai, L. Matthews, A. Osorio, S. Shanley, I. Giddings, F. Moreews, I. Locke, D.G. Evans, D. Eccles, R.D. Williams, M. Girolami, C. Campbell, and R. Eeles. Accurate prediction of brca1 and brca2 heterozygous genotype using expression profiling after induced dna damage. Clinical Cancer Research, 12(13):3896–3901, 2006.
M. Girolami, H. Mischak, and R. Krebs. Analysis of complex, multidimensional datasets. Drug Discovery Today: Technologies, 3(1):13–19, 2006.
A. Szymkowiak-Have, M.A. Girolami, and J. Larsen. Clustering via kernel decomposition. IEEE Transactions on Neural Networks, 17(1):256–264, 2006.
L. Carrivick, S. Rogers, J. Clark, C. Campbell, M. Girolami, and C. Cooper. Identification of prognostic signatures in breast cancer microarray data using bayesian techniques. Journal of the Royal Society Interface, 3(8):367–381, 2006.
M. Girolami and S. Rogers. Variational bayesian multinomial probit regression with gaussian process priors. Neural Computation, 18(8):1790–1817, 2006.
S. Rogers and M. Girolami. A bayesian regression approach to the inference of regulatory networks from gene expression data. Bioinformatics, 21(14):3131–3137, 2005.
S. Rogers, M. Girolami, R. Krebs, and H. Mischak. Disease classification from capillary electrophoresis: mass spectrometry. Lecture Notes in Computer Science, 3686(PART I):183–191, 2005.
M. Girolami and S. Rogers. Hierarchic bayesian models for kernel learning. ICML 2005 - Proceedings of the 22nd International Conference on Machine Learning, pages 241–248, 2005.
L. Azzopardi, M. Girolami, and M. Crowe. Probabilistic hyperspace analogue to language. SIGIR 2005 - Proceedings of the 28th Annual International ACM SIGIR Conference on Research and Development in Information Retrieval, pages 575–576, 2005.
M. Girolami and A. Kabán. Sequential activity profiling: latent dirichlet allocation of markov chains. Data Mining and Knowledge Discovery, 10(3):175–196, 2005.
S. Rogers, M. Girolami, C. Campbell, and R. Breitling. The latent process decomposition of cdna microarray data sets. IEEE/ACM Transactions on Computational Biology and Bioinformatics, 2(2):143–156, 2005.
A. Al-Shahib, C. He, A.C. Tan, M. Girolami, and D. Gilbert. An assessment of feature relevance in predicting protein function from sequence. Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), 3177:52–57, 2004.
M. Girolami and R. Breitling. Biologically valid linear factor models of gene expressions. Bioinformatics, 20(17):3021–3033, 2004.
C. He, M. Girolami, and G. Ross. Employing optimized combinations of one-class classifiers for automated currency validation. Pattern Recognition, 37(6):1085–1096, 2004.
C. He and M. Girolami. Novelty detection employing an l<inf>2</inf> optimal non-parametric density estimator. Pattern Recognition Letters, 25(12):1389–1397, 2004.
M. Girolami and A. Kabán. Simplicial mixtures of markov chains: distributed modelling of dynamic user profiles. Advances in Neural Information Processing Systems, 2004.
M. Girolami. Statistical and probabilistic fundamentals of ica. IET Seminar Digest, 2004(10774):3, 2004.
L. Azzopardi, M. Girolami, and C.J. Van Rijsbergen. Topic based language models for ad hoc information retrieval. IEEE International Conference on Neural Networks - Conference Proceedings, 4:3281–3286, 2004.
L. Azzopardi, M. Girolami, and C.J. Van Rijsbergen. User biased document language modelling. Proceedings of Sheffield SIGIR - Twenty-Seventh Annual International ACM SIGIR Conference on Research and Development in Information Retrieval, pages 542–543, 2004.
L. Azzopardi, M. Girolami, and K. Van Risjbergen. Investigating the relationship between language model perplexity and ir precision-recall measures. SIGIR Forum (ACM Special Interest Group on Information Retrieval), pages 369–370, 2003.
M. Girolami and A. Kabán. On an equivalence between plsi and lda. SIGIR Forum (ACM Special Interest Group on Information Retrieval), pages 433–434, 2003.
M. Girolami and C. He. Probability density estimation from optimally condensed data samples. IEEE Transactions on Pattern Analysis and Machine Intelligence, 25(10):1253–1264, 2003.
E. Bingham, A. Kabán, and M. Girolami. Topic identification in dynamical text by complexity pursuit. Neural Processing Letters, 17(1):69–83, 2003.
A. Kabán and M.A. Girolami. A dynamic probabilistic model to visualise topic evolution in text streams. Journal of Intelligent Information Systems, 18(2-3):107–125, 2002.
A. Kabán, P. Ti?o, and M. Girolami. A general framework for a principled hierarchical visualization of multivariate data. Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), 2412:518–523, 2002.
A. Vinokourov and M. Girolami. A probabilistic framework for the hierarchic organisation and classification of document collections. Journal of Intelligent Information Systems, 18(2-3):153–172, 2002.
D.R. Campbell, C. Fyfe, and M. Girolami. Artificial intelligence systems techniques and applications in speech processing. Intelligent Systems: Technology and Applications, Six Volume Set, pages III–1–III–48, 2002.
A. Kabán and M.A. Girolami. Fast extraction of semantic features from a latent semantic indexed text corpus. Neural Processing Letters, 15(1):31–43, 2002.
M. Girolami. Latent variable models for the topographic organisation of discrete and strictly positive data. Neurocomputing, 48(1-4):185–198, 2002.
M. Girolami. Mercer kernel-based clustering in feature space. IEEE Transactions on Neural Networks, 13(3):780–784, 2002.
M. Girolami. Orthogonal series density estimation and the kernel eigenvalue problem. Neural Computation, 14(3):669–688, 2002.
F. Crestani, M. Girolami, and K. van Rijsbergen. Preface. Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), 2291:V–VI, 2002.
F. Crestani, M. Girolami, and K. Van Rijsbergen. Report on the 24th european colloquium on information retrieval research (ecir 2002). SIGMOD Record, 31(3):77–80, 2002.
A. Kabán and M. Girolami. A combined latent class and trait model for the analysis and visualization of discrete data. IEEE Transactions on Pattern Analysis and Machine Intelligence, 23(8):859–872, 2001.
M. Girolami. A variational method for learning sparse and overcomplete representations. Neural Computation, 13(11):2517–2532, 2001.
R. Rosipal and M. Girolami. An expectation-maximization approach to nonlinear component analysis. Neural Computation, 13(3):505–510, 2001.
R. Rosipal, M. Girolami, L.J. Trejo, and A. Cichocki. Kernel pca for feature extraction and de-noising in nonlinear regression. Neural Computing and Applications, 10(3):231–243, 2001.
M. Girolami. The topographic organization and visualization of binary data using multivariate-bernoulli latent variable models. IEEE Transactions on Neural Networks, 12(6):1367–1374, 2001.
A. Vinokourov and M. Girolami. A probabilistic hierarchical clustering method for organising collections of text documents. Proceedings-International Conference on Pattern Recognition, 15(2):182–185, 2000.
T.-W. Lee, M. Girolami, A.J. Bell, and T.J. Sejnowski. A unifying information-theoretic framework for independent component analysis. Computers and Mathematics with Applications, 39(11):1–21, 2000.
A. Kabán and M. Girolami. Initialized and guided em-clustering of sparse binary data with application to text based documents. Proceedings - International Conference on Pattern Recognition, 15(2):744–747, 2000.
M. Girolami, A. Vinokourov, and A. Kabán. The organisation and visualisation of document corpora: a probabilistic approach. Proceedings - International Workshop on Database and Expert Systems Applications, DEXA, 2000-January:558–564, 2000.
Roman Rosipal and Mark Girolami. Adaptive support vector regression filter: a signal detection application. IEE Conference Publication, 2(470):603–607, 1999.
T.-W. Lee, M.S. Lewicki, M. Girolami, and T.J. Sejnowski. Blind source separation of more sources than mixtures using overcomplete representations. IEEE Signal Processing Letters, 6(4):87–90, 1999.
T.-W. Lee, M. Girolami, and T.J. Sejnowski. Independent component analysis using an extended infomax algorithm for mixed subgaussian and supergaussian sources. Neural Computation, 11(2):417–441, 1999.
M. Girolami, A. Cichocki, and S.-I. Amari. A common neural-network model for unsupervised exploratory data analysis and independent component analysis. IEEE Transactions on Neural Networks, 9(6):1495–1501, 1998.
M. Girolami. A nonlinear model of the binaural cocktail party effect. Neurocomputing, 22(1-3):201–215, 1998.
M. Girolami. An alternative perspective on adaptive independent component analysis algorithms. Neural Computation, 10(8):2103–2114, 1998.
M. Girolami. Noise reduction and speech enhancement via temporal anti-hebbian learning. ICASSP, IEEE International Conference on Acoustics, Speech and Signal Processing - Proceedings, 2:1233–1236, 1998.
M. Girolami. The latent variable data model for exploratory data analysis and visualisation: a generalisation of the nonlinear infomax algorithm. Neural Processing Letters, 8(1):27–39, 1998.
M. Girolami and C. Fyfe. An extended exploratory projection pursuit network with linear and nonlinear anti-hebbian lateral connections applied to the cocktail party problem. Neural Networks, 10(9):1607–1618, 1997.
M. Girolami and C. Fyfe. Extraction of independent signal sources using a deflationary exploratory projection pursuit network with lateral inhibition. IEE Proceedings: Vision, Image and Signal Processing, 144(5):299–306, 1997.
M. Girolami and C. Fyfe. Generalised independent component analysis through unsupervised learning with emergent bussgang properties. IEEE International Conference on Neural Networks - Conference Proceedings, 3:1788–1791, 1997.
Mark Girolami and Colin Fyfe. Kurtosis extrema and identification of independent components: a neural network approach. ICASSP, IEEE International Conference on Acoustics, Speech and Signal Processing - Proceedings, 4:3329–3332, 1997.
M. Girolami. Symmetric adaptive maximum likelihood estimation for noise cancellation and signal separation. Electronics Letters, 33(17):1437–1438, 1997.
M. Girolami and C. Fyfe. A temporal model of linear anti-hebbian learning. Neural Processing Letters, 4(3):139–148, 1996.